Code
library(duckplyr)
library(stringr)
library(dplyr)
library(arrow)
library(tidyr)
library(ggupset)
library(purrr)
library(forcats)
library(ggplot2)
Steven Turnbull
November 5, 2024
StatsNZ recently released their Aotearoa Data Explorer, a tool that makes accessing public data more accessible. The data is comprehensive and pretty clean, but it’s pretty large (35 million rows), and has some quirks that we may want to handle now to save us pain at a later date.
Our first obstacle is working with big data. Luckily, the duckdb makes it really easy. As this post outlines, we can use the duckplyr library in R to load in our big data, manipulate it using dplyr functions, and it will find the most efficient course through our code and return our clean data. duckplyr is a great package because it does what it can in the most efficient way using duckdb, and for anything else it will revert to dplyr as normal, meaning our code will run through. If you haven’t tried out duckdb yet, this method makes practically zero difference to your normal tidyverse workflow.
Here are the libraries we’ll need for running our code.
In our case, we want to reduce the size of the dataset by only including census year 2023, and removing a couple of categories with unknown values.
Multiple total count values are also included within each category, which can be useful in specific cases but is likely lead to some unforseen errors with double counting if we came back to use this dataset at a later date and forget it’s there. To make our future lives easier, I’ll exclude these cases - we can always get the total count by summing up the values in that category/combination of categories.
I’ll also use regex to add in a couple more columns that explain what each row represents in terms of the Area type and Age Band type. This will make it easier to work with in future.
time1 <- Sys.time()
data <- duckplyr_df_from_csv("STATSNZ,CEN23_ECI_008,1.0+all.csv") #Data downloaded from https://explore.data.stats.govt.nz/?fs[0]=2023%20Census%2C0%7CPopulation%20structure%23CAT_POPULATION_STRUCTURE%23&pg=0&fc=2023%20Census&bp=true&snb=20
df <- data |>
#select the columns we want
select(
`CEN23_YEAR_001: Census year`,
`CEN23_GEO_002: Area`,
`CEN23_ETH_006: Ethnicity`,
`CEN23_AGE_003: Age`,
`CEN23_GEN_002: Gender`,
OBS_VALUE
) |>
#give them nicer names
rename(
Year = `CEN23_YEAR_001: Census year`,
Area = `CEN23_GEO_002: Area`,
Ethnicity = `CEN23_ETH_006: Ethnicity`,
Age = `CEN23_AGE_003: Age`,
Gender = `CEN23_GEN_002: Gender`,
Value = OBS_VALUE
) |>
#remove rows with summary stats, we don't want to keep total values or medians.
filter(!Age %in% c('99: Total - age','Median: Median - age')) |>
#remove unknown gender
filter(!Gender %in% c(99)) |>
#just filter to 2023 census
filter(Year == "2023: 2023") |>
#remove missing ethnicities
filter(!str_detect(Ethnicity,"^\\d{4}")) |>
#assign an area type based on the area code
mutate(Area_Type = case_when(
str_detect(Area,"^9999") ~ "Total",
str_detect(Area,"^\\d{1}0:") ~ "Region",
str_detect(Area,"^\\d{2}:") ~ "Regional Council Area",
str_detect(Area,"^\\d{3}:") ~ "Territorial Authority",
str_detect(Area,"^\\d{5}:") ~ "Local Board Area",
str_detect(Area,"^\\d{6}:") ~ "SA22023",
TRUE ~ NA_character_
)) |>
#assign an age band type based on the age code
mutate(Age_Type = case_when(
str_detect(Age,"^\\d{1}:") ~ "15_year_bands",
str_detect(Age,"^\\d{2}:") ~ "5_year_bands",
TRUE ~ NA_character_
)) |>
# tidy
select(Year, Area_Type, Area, Age_Type, Age, Ethnicity, Gender, Value)
#save
df |>
write.csv("Population/census_population_2023_clean.csv")
#save as parquet (nicer to use later on)
df |>
write_parquet("Population/census_population_2023_clean.parquet")
cat("Time taken: ", Sys.time() - time1)The data we have from StatsNZ represents ethnicity in a way that maximises the amount of information about the groups people identify with using combinations of ethnicity groups. This is really cool, as it gives depth to our numbers and allows us to treat people how they want to be treated. When working with big data, it can be really easy to forget that each count represents someone significant, each value is a person, and behind each category is a choice that someone made.
Often categories such as ethnicity are simplified down as it can be tricky to summarise in a cohesive way. Maintaining some complexity in the categories we want to represent can introduce additional design decisions. With so many ethnicity groupings, axis labels can get cluttered and hard to read. Luckily, there is an easy way!
This is the perfect use case for what are known as UpSet Charts. While the name has depressing connotations, the plot itself will make everyone’s life easier. Simply put, an UpSet chart represents the combinations of a catgeorical variable as an additional plot where the axis text would normally sit. This walkthrough by Albert Rapp shows how we can make these plots pretty easily using the ggupset library in R.
But before we get to that, I’ll carry out some additional cleaning steps, and reshape our data to work with the ggupset library.
df_ethnicity <- df |>
#filter to one age band category to prevent double counting
filter(Age_Type == "15_year_bands") |>
# use total count
filter(Area == "9999: Total - New Zealand by regional council") |>
# summarise counts for each ethnicity
group_by(Ethnicity) |>
summarise(total = sum(Value,na.rm = T)) duckplyr: materializing, review details with duckplyr::last_rel()#and some extra cleaning steps
df_ethnicity_clean <- df_ethnicity |>
#tidy up some ethnicity labels (yes i love regex)
mutate(Ethnicity = str_replace_all(Ethnicity, "Middle Eastern/Latin American/African","MELAA")) |>
mutate(Ethnicity = str_replace_all(Ethnicity, ".*\\: ","")) |>
mutate(Ethnicity = str_replace_all(Ethnicity, " only",""))
# get a TRUE or FALSE value for each category showing whether an ethnicity category applies
ggupset_df <- df_ethnicity_clean |>
mutate(European = ifelse(str_detect(Ethnicity, "European"),T,F))|>
mutate(Māori = ifelse(str_detect(Ethnicity, "Māori"),T,F))|>
mutate(`Pacific Peoples` = ifelse(str_detect(Ethnicity, "Pacific Peoples"),T,F))|>
mutate(Asian = ifelse(str_detect(Ethnicity, "Asian"),T,F))|>
mutate(MELAA = ifelse(str_detect(Ethnicity, "MELAA"),T,F))|>
mutate(`Other Ethnicity` = ifelse(str_detect(Ethnicity, "Other Ethnicity"),T,F))|>
# add column for combination values as list for each row
mutate(
Combination = pmap(
list(European, Māori, `Pacific Peoples`, Asian, MELAA, `Other Ethnicity`),
\(lgl1, lgl2, lgl3, lgl4, lgl5, lgl6) {
c('European', 'Māori', 'Pacific Peoples', 'Asian', 'MELAA', 'Other Ethnicity')[c(lgl1, lgl2, lgl3, lgl4, lgl5, lgl6)]
}
)
) And plot!
# Plot using ggtern
ggupset_df|>
#just show the top 15 combinations by total population
head(15) |>
ggplot(aes(x = Combination,y = total)) +
geom_bar(stat="identity") +
# add the count to the bar as a tidy label
geom_text(aes(label = scales::comma(total)), vjust = -0.5, size=2) +
# add the combinations as the x axis
scale_x_upset(order_by = "degree") +
#tidy our y axis values
scale_y_continuous(labels = scales::comma) +
#tidy up lavels
labs(
title = "Ethnicity Group Distribution (Census 2023)",
x = "",
y = "Total Population",
caption = str_wrap("https://explore.data.stats.govt.nz/ : Ethnicity (detailed single / combination), age, and gender for the census usually resident population count, (RC, TALB, SA2, Health), 2013, 2018, and 2023 Censuses. MELAA = Middle Eastern, Latin American and African.",100)
) +
# update the UpSet plot to have red dots, because why not
theme_combmatrix(
combmatrix.panel.point.color.fill = "red"
)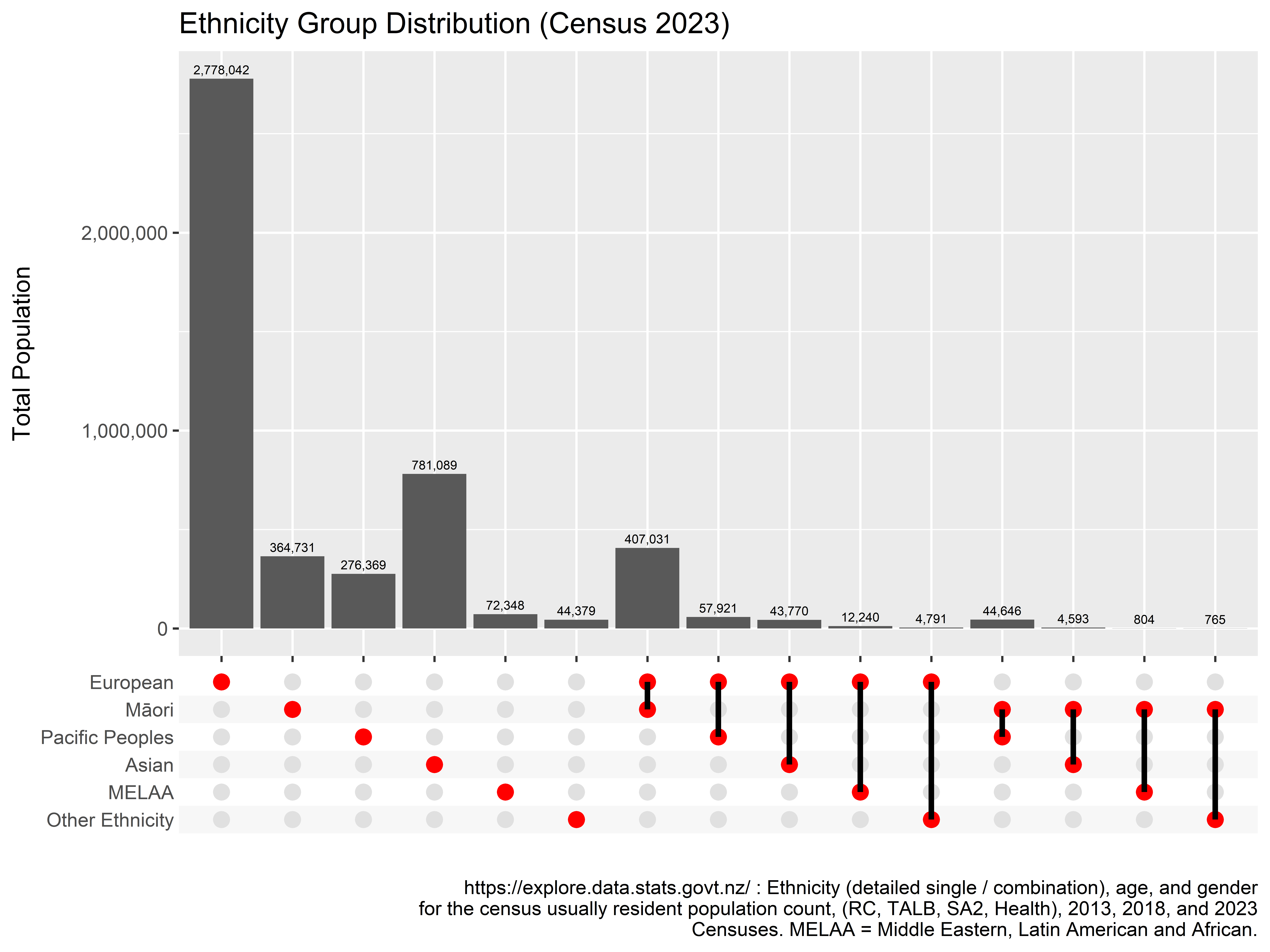
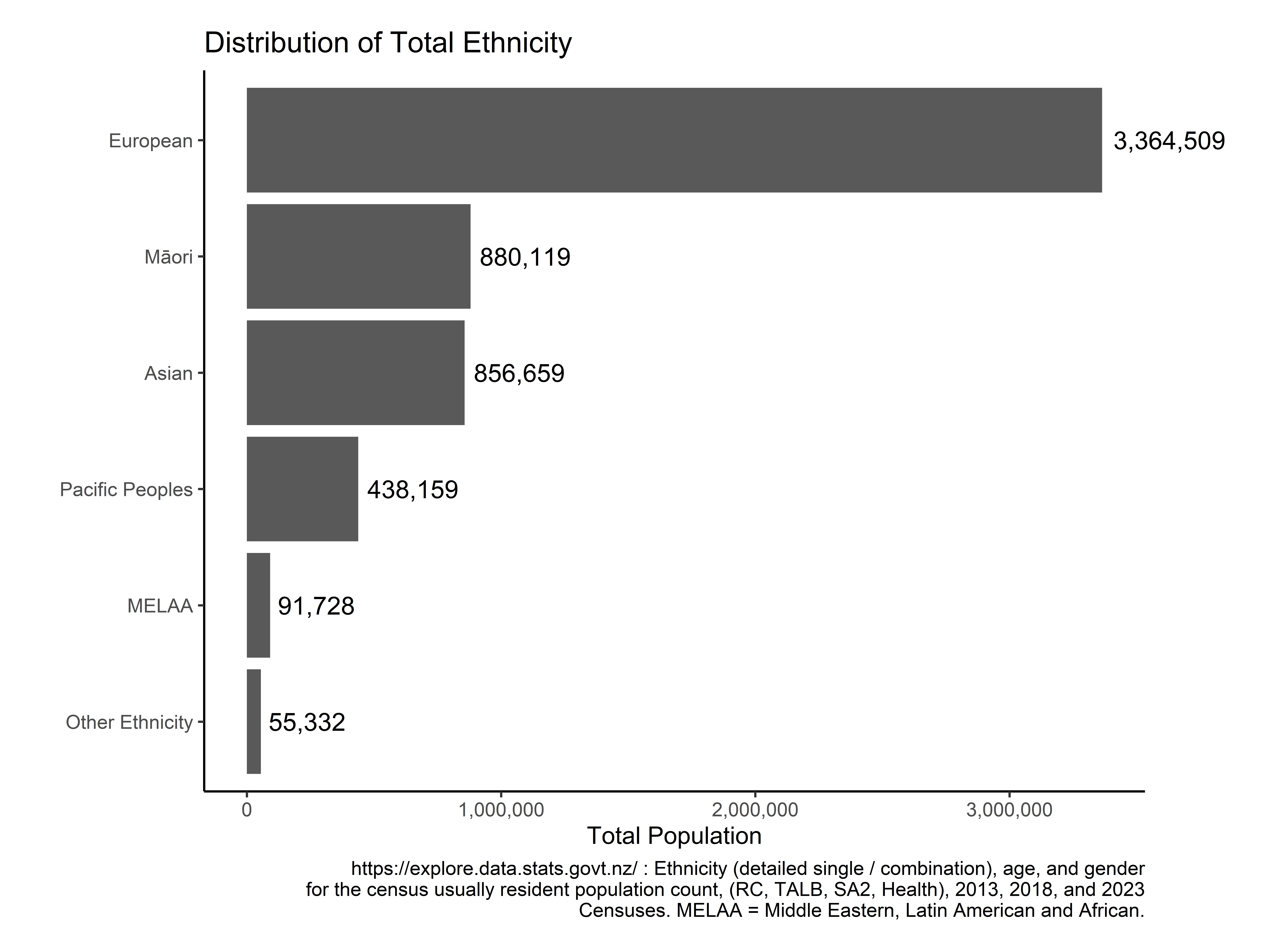
UpSet plots make it really easy to identify which bars represent which categories, and when you have lots of combinations this is needed. One key thing that stands out in our data is that the Asian Only category is the second highest behind European. At first glance this may seem incorrect, we might expect Māori to be higher. However, plotting the total counts across categories using each combination (i.e, totalling up Māori, Māori/European, Māori/Pacific etc. into one group labelled Māori), we can see that Māori represent the second highest population overall. It just happens that Māori are more likely to identify with an additional ethnicity, while Asian are more likely to identify as only Asian.
#get count for each category overall
unnest(ggupset_df, Combination) |>
group_by(Combination) |>
summarise(Total = sum(total)) |>
#plot
ggplot(aes(y = fct_reorder(Combination,Total), x = Total)) +
geom_bar(stat = "identity") +
geom_text(
aes(label = scales::comma(Total)),
hjust = -0.1, size = 4
) +
scale_x_continuous(labels = scales::comma) +
labs(
title = "Distribution of Total Ethnicity",
y = "",
x = "Total Population",
caption = str_wrap("https://explore.data.stats.govt.nz/ : Ethnicity (detailed single / combination), age, and gender for the census usually resident population count, (RC, TALB, SA2, Health), 2013, 2018, and 2023 Censuses. MELAA = Middle Eastern, Latin American and African.",100)
) +
theme_classic() +
#stop the labels getting clipped off
coord_cartesian(clip="off") +
theme(
plot.margin = margin(0.5,2,0.5,0.5,"cm")
)
Being able to easily visualise and explore our data in depth provides significant insights that we can miss when prioritising simpler, less detailed views of data.
---
title: 'Visualising Ethnicity from NZ Census 2023'
date: '2024-11-05'
categories: ['R', 'Population','Visualisation','New Zealand']
description: 'In this post, I talk about why you should treat big data as ducks 🦆, see populations as people 👨👩👧👦, and why UpSet plots will make you happy 😃.'
author: 'Steven Turnbull'
execute:
message: false
warning: false
editor_options:
chunk_output_type: console
format:
html:
code-fold: show
code-tools: true
---
# Big Data as Ducks(db) 🦆
StatsNZ recently released their [Aotearoa Data Explorer](https://www.stats.govt.nz/tools/aotearoa-data-explorer/), a tool that makes accessing public data more accessible. The data is comprehensive and pretty clean, but it's pretty large (35 million rows), and has some quirks that we may want to handle now to save us pain at a later date.
Our first obstacle is working with big data. Luckily, the `duckdb` makes it really easy. As [this post](https://duckdb.org/2024/10/09/analyzing-open-government-data-with-duckplyr) outlines, we can use the `duckplyr` library in R to load in our big data, manipulate it using dplyr functions, and it will find the most efficient course through our code and return our clean data. `duckplyr` is a great package because it does what it can in the most efficient way using duckdb, and for anything else it will revert to dplyr as normal, meaning our code will run through. If you haven't tried out duckdb yet, this method makes practically zero difference to your normal tidyverse workflow.
Here are the libraries we'll need for running our code.
```{r, message = F, warning = F}
library(duckplyr)
library(stringr)
library(dplyr)
library(arrow)
library(tidyr)
library(ggupset)
library(purrr)
library(forcats)
library(ggplot2)
```
In our case, we want to reduce the size of the dataset by only including census year 2023, and removing a couple of categories with unknown values.
Multiple total count values are also included within each category, which can be useful in specific cases but is likely lead to some unforseen errors with double counting if we came back to use this dataset at a later date and forget it's there. To make our future lives easier, I'll exclude these cases - we can always get the total count by summing up the values in that category/combination of categories.
I'll also use [regex](https://r4ds.hadley.nz/regexps) to add in a couple more columns that explain what each row represents in terms of the Area type and Age Band type. This will make it easier to work with in future.
```{r, eval = F, message = F, warning = F}
time1 <- Sys.time()
data <- duckplyr_df_from_csv("STATSNZ,CEN23_ECI_008,1.0+all.csv") #Data downloaded from https://explore.data.stats.govt.nz/?fs[0]=2023%20Census%2C0%7CPopulation%20structure%23CAT_POPULATION_STRUCTURE%23&pg=0&fc=2023%20Census&bp=true&snb=20
df <- data |>
#select the columns we want
select(
`CEN23_YEAR_001: Census year`,
`CEN23_GEO_002: Area`,
`CEN23_ETH_006: Ethnicity`,
`CEN23_AGE_003: Age`,
`CEN23_GEN_002: Gender`,
OBS_VALUE
) |>
#give them nicer names
rename(
Year = `CEN23_YEAR_001: Census year`,
Area = `CEN23_GEO_002: Area`,
Ethnicity = `CEN23_ETH_006: Ethnicity`,
Age = `CEN23_AGE_003: Age`,
Gender = `CEN23_GEN_002: Gender`,
Value = OBS_VALUE
) |>
#remove rows with summary stats, we don't want to keep total values or medians.
filter(!Age %in% c('99: Total - age','Median: Median - age')) |>
#remove unknown gender
filter(!Gender %in% c(99)) |>
#just filter to 2023 census
filter(Year == "2023: 2023") |>
#remove missing ethnicities
filter(!str_detect(Ethnicity,"^\\d{4}")) |>
#assign an area type based on the area code
mutate(Area_Type = case_when(
str_detect(Area,"^9999") ~ "Total",
str_detect(Area,"^\\d{1}0:") ~ "Region",
str_detect(Area,"^\\d{2}:") ~ "Regional Council Area",
str_detect(Area,"^\\d{3}:") ~ "Territorial Authority",
str_detect(Area,"^\\d{5}:") ~ "Local Board Area",
str_detect(Area,"^\\d{6}:") ~ "SA22023",
TRUE ~ NA_character_
)) |>
#assign an age band type based on the age code
mutate(Age_Type = case_when(
str_detect(Age,"^\\d{1}:") ~ "15_year_bands",
str_detect(Age,"^\\d{2}:") ~ "5_year_bands",
TRUE ~ NA_character_
)) |>
# tidy
select(Year, Area_Type, Area, Age_Type, Age, Ethnicity, Gender, Value)
#save
df |>
write.csv("Population/census_population_2023_clean.csv")
#save as parquet (nicer to use later on)
df |>
write_parquet("Population/census_population_2023_clean.parquet")
cat("Time taken: ", Sys.time() - time1)
```
# Treating Populations as People 👨👩👧👦
The data we have from StatsNZ represents ethnicity in a way that maximises the amount of information about the groups people identify with using combinations of ethnicity groups. This is really cool, as it gives depth to our numbers and allows us to treat people how they want to be treated. When working with big data, it can be really easy to forget that each count represents someone significant, each value is a person, and behind each category is a choice that someone made.
Often categories such as ethnicity are simplified down as it can be tricky to summarise in a cohesive way. Maintaining some complexity in the categories we want to represent can introduce additional design decisions. With so many ethnicity groupings, axis labels can get cluttered and hard to read. Luckily, there is an easy way!
```{r, include = F}
df <- read_parquet("C:/Users/steven.turnbull_syne/Documents/GitHub/sturbull.github.io/inputs/Population/census_population_2023_clean.parquet")
```
# UpSet Plots Will Make You Happy 😃
This is the perfect use case for what are known as `UpSet Charts`. While the name has depressing connotations, the plot itself will make everyone's life easier. Simply put, an UpSet chart represents the combinations of a catgeorical variable as an additional plot where the axis text would normally sit. [This walkthrough by Albert Rapp](https://albert-rapp.de/posts/ggplot2-tips/26_upset_charts/26_upset_charts.html) shows how we can make these plots pretty easily using the `ggupset` library in R.
But before we get to that, I'll carry out some additional cleaning steps, and reshape our data to work with the `ggupset` library.
```{r,message=F,warning=F}
df_ethnicity <- df |>
#filter to one age band category to prevent double counting
filter(Age_Type == "15_year_bands") |>
# use total count
filter(Area == "9999: Total - New Zealand by regional council") |>
# summarise counts for each ethnicity
group_by(Ethnicity) |>
summarise(total = sum(Value,na.rm = T))
#and some extra cleaning steps
df_ethnicity_clean <- df_ethnicity |>
#tidy up some ethnicity labels (yes i love regex)
mutate(Ethnicity = str_replace_all(Ethnicity, "Middle Eastern/Latin American/African","MELAA")) |>
mutate(Ethnicity = str_replace_all(Ethnicity, ".*\\: ","")) |>
mutate(Ethnicity = str_replace_all(Ethnicity, " only",""))
# get a TRUE or FALSE value for each category showing whether an ethnicity category applies
ggupset_df <- df_ethnicity_clean |>
mutate(European = ifelse(str_detect(Ethnicity, "European"),T,F))|>
mutate(Māori = ifelse(str_detect(Ethnicity, "Māori"),T,F))|>
mutate(`Pacific Peoples` = ifelse(str_detect(Ethnicity, "Pacific Peoples"),T,F))|>
mutate(Asian = ifelse(str_detect(Ethnicity, "Asian"),T,F))|>
mutate(MELAA = ifelse(str_detect(Ethnicity, "MELAA"),T,F))|>
mutate(`Other Ethnicity` = ifelse(str_detect(Ethnicity, "Other Ethnicity"),T,F))|>
# add column for combination values as list for each row
mutate(
Combination = pmap(
list(European, Māori, `Pacific Peoples`, Asian, MELAA, `Other Ethnicity`),
\(lgl1, lgl2, lgl3, lgl4, lgl5, lgl6) {
c('European', 'Māori', 'Pacific Peoples', 'Asian', 'MELAA', 'Other Ethnicity')[c(lgl1, lgl2, lgl3, lgl4, lgl5, lgl6)]
}
)
)
```
And plot!
```{r,message=F,warning=F, fig.width=8,fig.height = 6,dpi=600}
# Plot using ggtern
ggupset_df|>
#just show the top 15 combinations by total population
head(15) |>
ggplot(aes(x = Combination,y = total)) +
geom_bar(stat="identity") +
# add the count to the bar as a tidy label
geom_text(aes(label = scales::comma(total)), vjust = -0.5, size=2) +
# add the combinations as the x axis
scale_x_upset(order_by = "degree") +
#tidy our y axis values
scale_y_continuous(labels = scales::comma) +
#tidy up lavels
labs(
title = "Ethnicity Group Distribution (Census 2023)",
x = "",
y = "Total Population",
caption = str_wrap("https://explore.data.stats.govt.nz/ : Ethnicity (detailed single / combination), age, and gender for the census usually resident population count, (RC, TALB, SA2, Health), 2013, 2018, and 2023 Censuses. MELAA = Middle Eastern, Latin American and African.",100)
) +
# update the UpSet plot to have red dots, because why not
theme_combmatrix(
combmatrix.panel.point.color.fill = "red"
)
```
UpSet plots make it really easy to identify which bars represent which categories, and when you have lots of combinations this is needed. One key thing that stands out in our data is that the Asian Only category is the second highest behind European. At first glance this may seem incorrect, we might expect Māori to be higher. However, plotting the total counts across categories using each combination (i.e, totalling up Māori, Māori/European, Māori/Pacific etc. into one group labelled Māori), we can see that Māori represent the second highest population overall. It just happens that Māori are more likely to identify with an additional ethnicity, while Asian are more likely to identify as only Asian.
```{r, message=F,warning=F, fig.width=8,fig.height = 6,dpi=600}
#get count for each category overall
unnest(ggupset_df, Combination) |>
group_by(Combination) |>
summarise(Total = sum(total)) |>
#plot
ggplot(aes(y = fct_reorder(Combination,Total), x = Total)) +
geom_bar(stat = "identity") +
geom_text(
aes(label = scales::comma(Total)),
hjust = -0.1, size = 4
) +
scale_x_continuous(labels = scales::comma) +
labs(
title = "Distribution of Total Ethnicity",
y = "",
x = "Total Population",
caption = str_wrap("https://explore.data.stats.govt.nz/ : Ethnicity (detailed single / combination), age, and gender for the census usually resident population count, (RC, TALB, SA2, Health), 2013, 2018, and 2023 Censuses. MELAA = Middle Eastern, Latin American and African.",100)
) +
theme_classic() +
#stop the labels getting clipped off
coord_cartesian(clip="off") +
theme(
plot.margin = margin(0.5,2,0.5,0.5,"cm")
)
```
Being able to easily visualise and explore our data in depth provides significant insights that we can miss when prioritising simpler, less detailed views of data.